Optimus Protein
New research has identified the RNA-binding protein DHX29 as a critical sensor that allows human cells to detect non-optimal codon usage and selectively regulate mRNA stability. Published in Science, the findings illuminate a long-standing mystery in molecular biology—how cells interpret subtle variations in the genetic code to fine-tune gene expression—potentially opening avenues for understanding disorders linked to dysregulated translation, such as certain cancers and inflammatory conditions.
The Hidden Layer in the Genetic Code
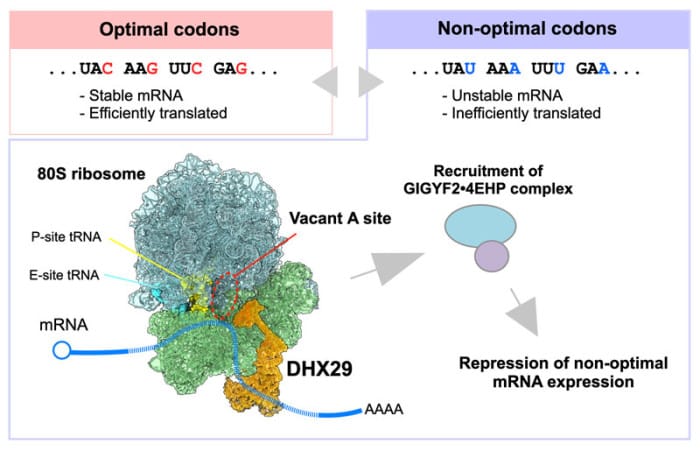
Genes are spelled out in codons—triplets of nucleotides that specify amino acids during protein synthesis. While multiple codons can encode the same amino acid (making them synonymous), they are not functionally identical. Some codons support efficient translation and stable mRNAs; others, termed non-optimal, slow translation and promote mRNA decay. This “hidden layer” of information influences global gene expression across species, yet the molecular machinery that detects non-optimal codons in human cells had remained elusive.
Pinpointing DHX29 Through Genome-Wide Screening
A collaborative team led by researchers at Kyoto University, including Osamu Takeuchi, Takuhiro Ito, and Masanori Yoshinaga, employed genome-wide CRISPR screening to hunt for factors that govern codon-dependent gene expression. The screen spotlighted DHX29, an RNA helicase previously known for its roles in translation initiation.
Results indicated that loss of DHX29 led to the upregulation of mRNAs enriched in non-optimal codons, confirming its role as a selective repressor. Further experiments showed that DHX29 preferentially associates with ribosomes actively decoding these less-favored codons.
A Direct Ribosomal Sensor and Recruitment Mechanism
Advanced structural and profiling techniques revealed how DHX29 operates. Cryo-electron microscopy demonstrated that DHX29 binds directly to the A-site entrance of the translating 80S ribosome—the precise location where the eEF1A•GTP•aminoacyl-tRNA complex delivers the next amino acid. This positioning allows DHX29 to monitor codon optimality during translation.
Selective ribosome profiling reinforced the finding: DHX29-bound ribosomes were enriched for non-optimal codons. Once engaged, DHX29 recruits the GIGYF2•4EHP protein complex, which in turn suppresses the targeted mRNAs, marking them for decay or translational repression.
The mechanism thus links codon choice directly to mRNA stability control, providing the first detailed picture of this process in human cells.
Evidence from Complementary Approaches
The study combined multiple orthogonal methods for robustness:
- RNA sequencing to quantify global shifts in mRNA levels upon DHX29 depletion.
- Proteomic analysis to identify the GIGYF2•4EHP interaction partners.
- Cryo-EM structures and ribosome profiling to map molecular interactions at near-atomic resolution.
These data collectively establish DHX29 as a ribosome-associated sentinel that translates codon bias into gene-expression outcomes.
Team leader Osamu Takeuchi reflected on the discovery: “We have long been fascinated by how cells interpret the hidden layer of information embedded within the genetic code, so discovering the molecular factor that allows human cells to read and respond to this hidden code has been particularly rewarding.”
Co-corresponding author Masanori Yoshinaga added: “Together, these findings reveal a direct molecular link between synonymous codon choice and the control of gene expression in human cells.”
Broader Implications for Cellular Health
The DHX29 pathway may influence fundamental processes such as cell differentiation, maintenance of cellular homeostasis, and responses to stress. Because codon optimality and mRNA stability are frequently altered in diseases—including cancer, where translation control is often hijacked—the work offers a new lens for investigating these conditions. While the study itself was conducted in human cell models, it lays groundwork for exploring DHX29’s contributions in more complex physiological contexts.
Toward Deeper Understanding and Future Applications
Although the results provide compelling mechanistic insight, the authors note that additional research will be required to determine how DHX29-mediated regulation operates in living tissues and whether it can be modulated for therapeutic benefit. The team plans to investigate the pathway’s roles in health and disease progression.
This discovery exemplifies how fundamental research continues to uncover elegant layers of control within the genome, offering measured progress toward a fuller picture of human gene regulation. As scientists probe these mechanisms further, the findings underscore the value of precise molecular tools in revealing biology’s hidden codes—step by step, with evidence guiding each advance.