Artificial Building Blocks of Life
DNA carries the genetic information of all living organisms and consists of only four different building blocks, the nucleotides. Nucleotides are composed of three distinctive parts: a sugar molecule, a phosphate group and one of the four nucleobases adenine, thymine, guanine and cytosine. The nucleotides are lined up millions of times and form the DNA double helix, similar to a spiral staircase. Scientists from the UoC’s Department of Chemistry have now shown that the structure of nucleotides can be modified to a great extent in the laboratory. The researchers developed so-called threofuranosyl nucleic acid (TNA) with a new, additional base pair. These are the first steps on the way to fully artificial nucleic acids with enhanced chemical functionalities. The study ‘Expanding the Horizon of the Xeno Nucleic Acid Space: Threose Nucleic Acids with Increased Information Storage’ was published in the Journal of the American Chemical Society.
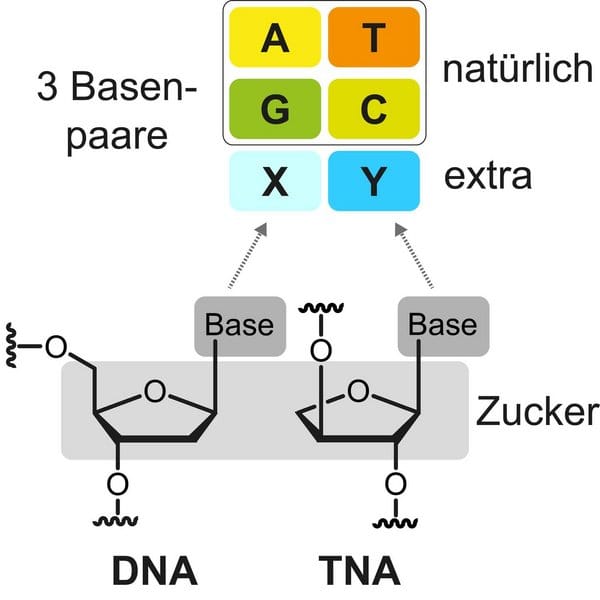
Artificial nucleic acids differ in structure from their originals. These changes affect their stability and function. “Our threofuranosyl nucleic acid is more stable than the naturally occurring nucleic acids DNA and RNA, which brings many advantages for future therapeutic use,” said Professor Dr Stephanie Kath-Schorr. For the study, the 5-carbon sugar deoxyribose, which forms the backbone in DNA, was replaced by a 4-carbon sugar. In addition, the number of nucleobases was increased from four to six. By exchanging the sugar, the TNA is not recognized by the cell's own degradation enzymes. This has been a problem with nucleic acid-based therapeutics, as synthetically produced RNA that is introduced into a cell is rapidly degraded and loses its effect. The introduction of TNAs into cells that remain undetected could now maintain the effect for longer.
“In addition, the built-in unnatural base pair enables alternative binding options to target molecules in the cell,” added Hannah Depmeier, lead author of the study. Kath-Schorr is certain that such a function can be used in particular in the development of new aptamers, short DNA or RNA sequences, which can be used for the targeted control of cellular mechanisms. TNAs could also be used for the targeted transport of drugs to specific organs in the body (targeted drug delivery) as well as in diagnostics; they could also be useful for the recognition of viral proteins or biomarkers.
Written by University of Cologne
Image by Dapmeier et al. & UoC
